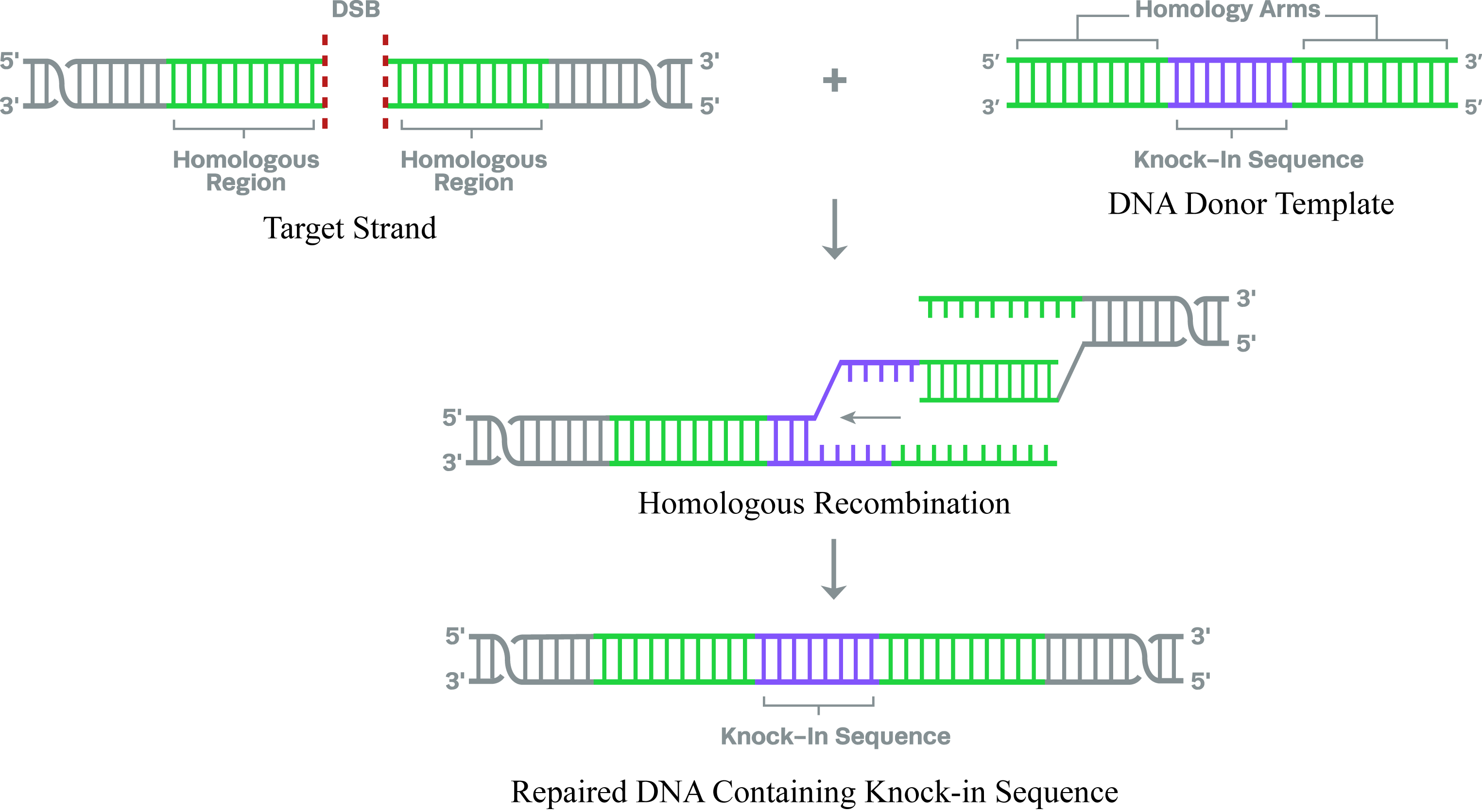
Design of CRISPR Knock-ins guide RNAs
CRISPR Knock-in experiments, wherein researchers insert a gene of interest at the specific site, rely on HDR. This mechanism uses a homologous template to repair DSBs and is therefore highly accurate. The precision of the HDR repair pathway can be coupled with the specificity of CRISPR-Cas to introduce the desired sequence into the target genomic region.More ....
Optimizing Success With CRISPR Knock-in Experiments:
- Choose the Right Guide RNA
- Pick the Right DNA Donor Template
- Single-stranded DNA (ssDNA)
- Avoid Re-cutting by Cas9
- Optimizing HDR efficiency over NHEJ