Plasmids used in our labs
CRISPR/Cas9
Some quick example text to build on the card title and make up the bulk of the card's content.
Read MoreGhCBE
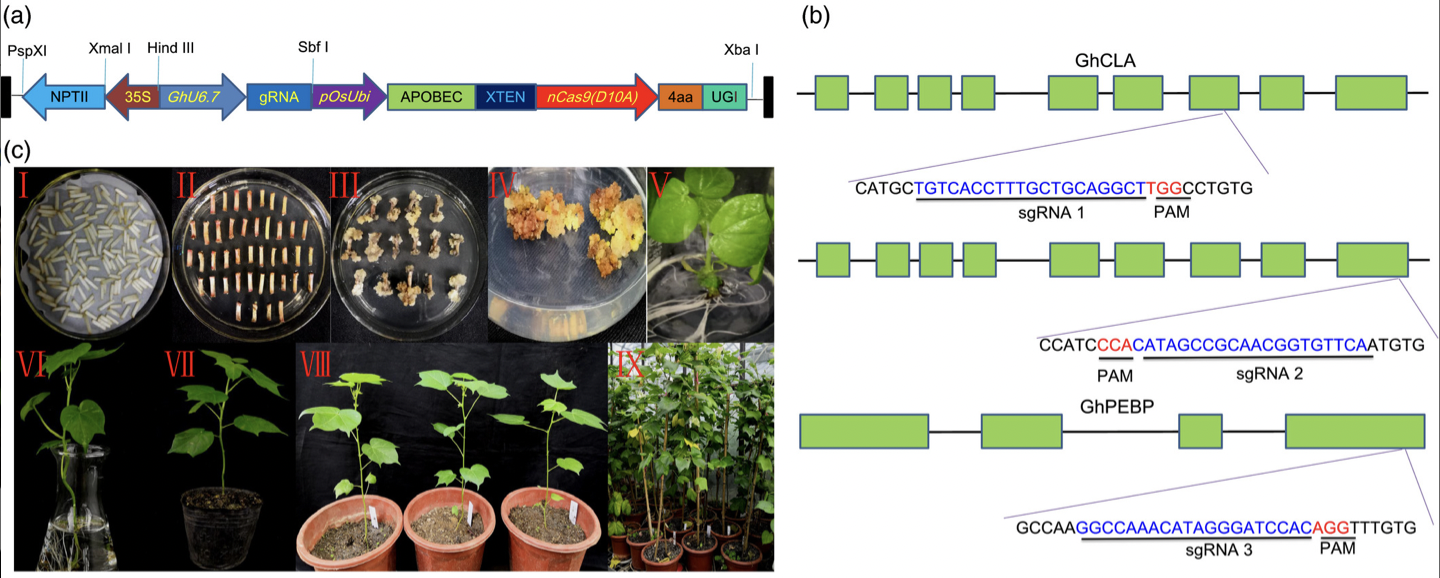
A cytidine deaminase sequence (APOBEC) fused with nCas9 and uracil glycosylase inhibitor (UGI) was inserted into our CRISPR/Cas9 plasmid (pRGEB32-GhU6.7). The editing efficiency ranged from 26.67 to 57.78% at the three target sites.
Read MoreGhABE
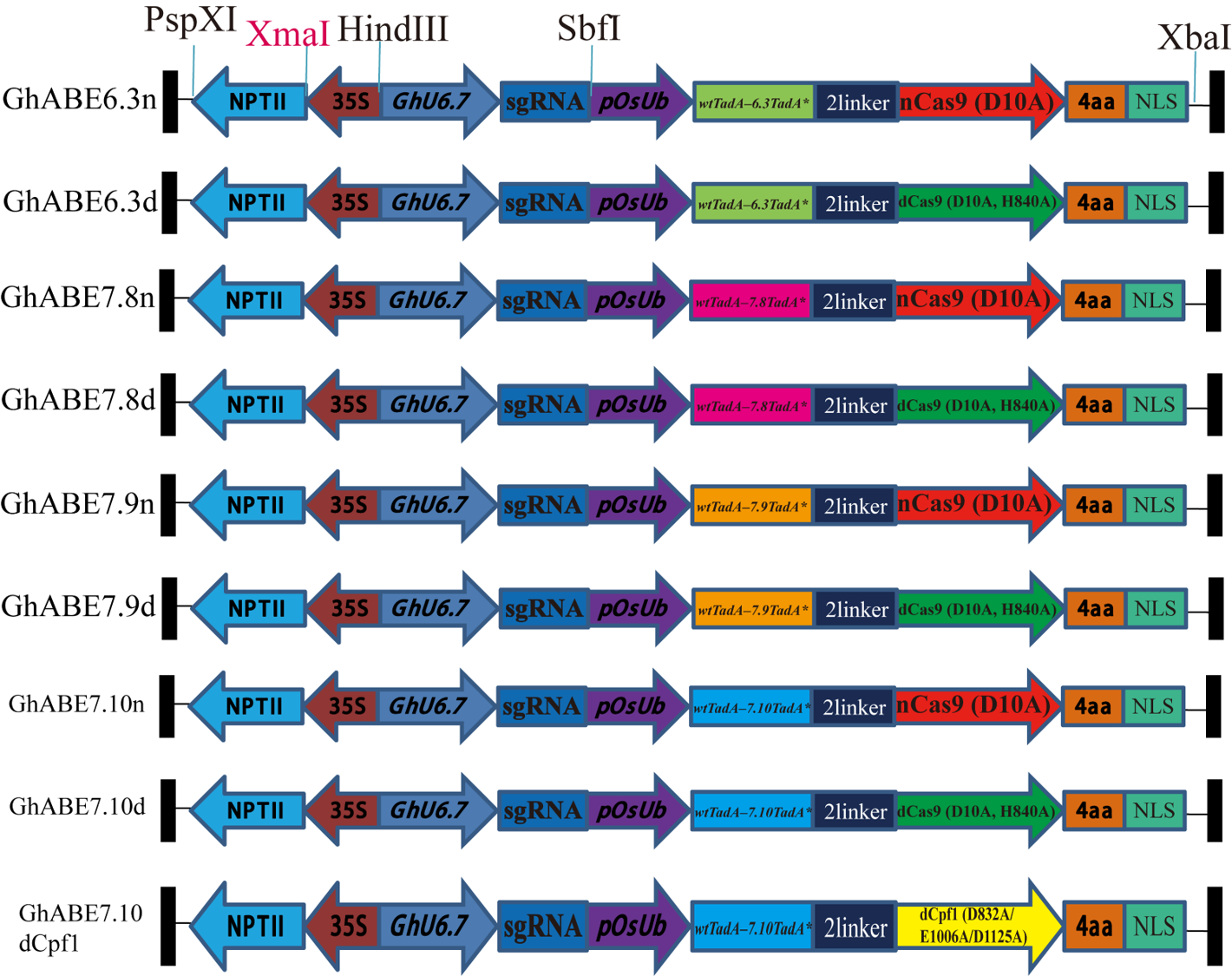
We established various ABE vectors based on different engineered adenosine deaminase (TadA) proteins fused to Cas9 variants (dCas9, nCas9), enabling efficient A to G editing up to 64% efficiency on-target sites of the allotetraploid cotton genome.
Read MoreGhCBE
A cytidine deaminase sequence (APOBEC) fused with nCas9 and uracil glycosylase inhibitor (UGI) was inserted into our CRISPR/Cas9 plasmid (pRGEB32-GhU6.7). The editing efficiency ranged from 26.67 to 57.78% at the three target sites.
Read MoreGhABE
We established various ABE vectors based on different engineered adenosine deaminase (TadA) proteins fused to Cas9 variants (dCas9, nCas9), enabling efficient A to G editing up to 64% efficiency on-target sites of the allotetraploid cotton genome.
Read MoreGhCBE + GhABE
Some quick example text to build on the card title and make up the bulk of the card's content.
Read MoreCRISPR-Cas12a (Cpf1)
A cytidine deaminase sequence (APOBEC) fused with nCas9 and uracil glycosylase inhibitor (UGI) was inserted into our CRISPR/Cas9 plasmid (pRGEB32-GhU6.7). The editing efficiency ranged from 26.67 to 57.78% at the three target sites.
Read MoreCRISPR-Cas12b (C2c1)
We established various ABE vectors based on different engineered adenosine deaminase (TadA) proteins fused to Cas9 variants (dCas9, nCas9), enabling efficient A to G editing up to 64% efficiency on-target sites of the allotetraploid cotton genome.
Read MoreCRISPR Cas13
Some quick example text to build on the card title and make up the bulk of the card's content.
Read MorePrime Editing
A cytidine deaminase sequence (APOBEC) fused with nCas9 and uracil glycosylase inhibitor (UGI) was inserted into our CRISPR/Cas9 plasmid (pRGEB32-GhU6.7). The editing efficiency ranged from 26.67 to 57.78% at the three target sites.
Read MoreCRISPR activation
We established various ABE vectors based on different engineered adenosine deaminase (TadA) proteins fused to Cas9 variants (dCas9, nCas9), enabling efficient A to G editing up to 64% efficiency on-target sites of the allotetraploid cotton genome.
Read MoreCRISPR Knock-ins
Some quick example text to build on the card title and make up the bulk of the card's content.
Read MoreCRISPR Epigenetics
A cytidine deaminase sequence (APOBEC) fused with nCas9 and uracil glycosylase inhibitor (UGI) was inserted into our CRISPR/Cas9 plasmid (pRGEB32-GhU6.7). The editing efficiency ranged from 26.67 to 57.78% at the three target sites.
Read MoreCRISPR activation
We established various ABE vectors based on different engineered adenosine deaminase (TadA) proteins fused to Cas9 variants (dCas9, nCas9), enabling efficient A to G editing up to 64% efficiency on-target sites of the allotetraploid cotton genome.
Read MoreCRISPR Knock-ins
Some quick example text to build on the card title and make up the bulk of the card's content.
Read More